This function generates a so-called diamond plot: a plot based on the forest plots that are commonplace in meta-analyses. The underlying idea is that point estimates are uninformative, and it would be better to focus on confidence intervals. The problem of the points with errorbars that are commonly employed is that the focus the audience's attention on the upper and lower bounds, even though those are the least relevant values. Using diamonds remedies this.
meansDiamondPlot(
data,
items = NULL,
labels = NULL,
decreasing = NULL,
conf.level = 0.95,
showData = TRUE,
dataAlpha = 0.1,
dataSize = 3,
dataColor = "#444444",
diamondColors = NULL,
jitterWidth = 0.5,
jitterHeight = 0.4,
returnLayerOnly = FALSE,
xlab = "Scores and means",
ylab = NULL,
theme = ggplot2::theme_bw(),
xbreaks = "auto",
outputFile = NULL,
outputWidth = 10,
outputHeight = 10,
ggsaveParams = ufs::opts$get("ggsaveParams"),
dat = NULL,
...
)Arguments
- data, dat
The dataframe containing the variables (
items) to show in the diamond plot (the namedatfor this argument is deprecated but still works for backward compatibility).- items
Optionally, the names (or numeric indices) of the variables (items) to show in the diamond plot. If NULL, all columns (variables, items) will be used.
- labels
A character vector of labels to use instead of column names from the dataframe.
- decreasing
Whether to sort the variables (rows) in the diamond plot decreasing (TRUE), increasing (FALSE), or not at all (NULL).
- conf.level
The confidence of the confidence intervals.
- showData
Whether to show the raw data or not.
- dataAlpha
This determines the alpha (transparency) of the data points. Note that argument
alphacan be used to set the alpha of the diamonds; this is eventually passed on toggDiamondLayer().- dataSize
The size of the data points.
- dataColor
The color of the data points.
- diamondColors
A vector of the same length as there are rows in the dataframe, to manually specify colors for the diamonds.
- jitterWidth
How much to jitter the individual datapoints horizontally.
- jitterHeight
How much to jitter the individual datapoints vertically.
- returnLayerOnly
Set this to TRUE to only return the
ggplot()layer of the diamondplot, which can be useful to include it in other plots.- xlab, ylab
The labels of the X and Y axes.
- theme
The theme to use.
- xbreaks
Where the breaks (major grid lines, ticks, and labels) on the x axis should be.
- outputFile
A file to which to save the plot.
- outputWidth, outputHeight
Width and height of saved plot (specified in centimeters by default, see
ggsaveParams).- ggsaveParams
Parameters to pass to ggsave when saving the plot.
- ...
Additional arguments are passed to
diamondPlot()and eventually toggDiamondLayer(). This can be used to, for example, specify two or more colors to use to generate a gradient (usinggenerateColorsand maybefullColorRange).
Value
A ggplot() plot with a ggDiamondLayer() is
returned.
Examples
tmpDf <- data.frame(item1 = rnorm(50, 1.6, 1),
item2 = rnorm(50, 2.6, 2),
item3 = rnorm(50, 4.1, 3));
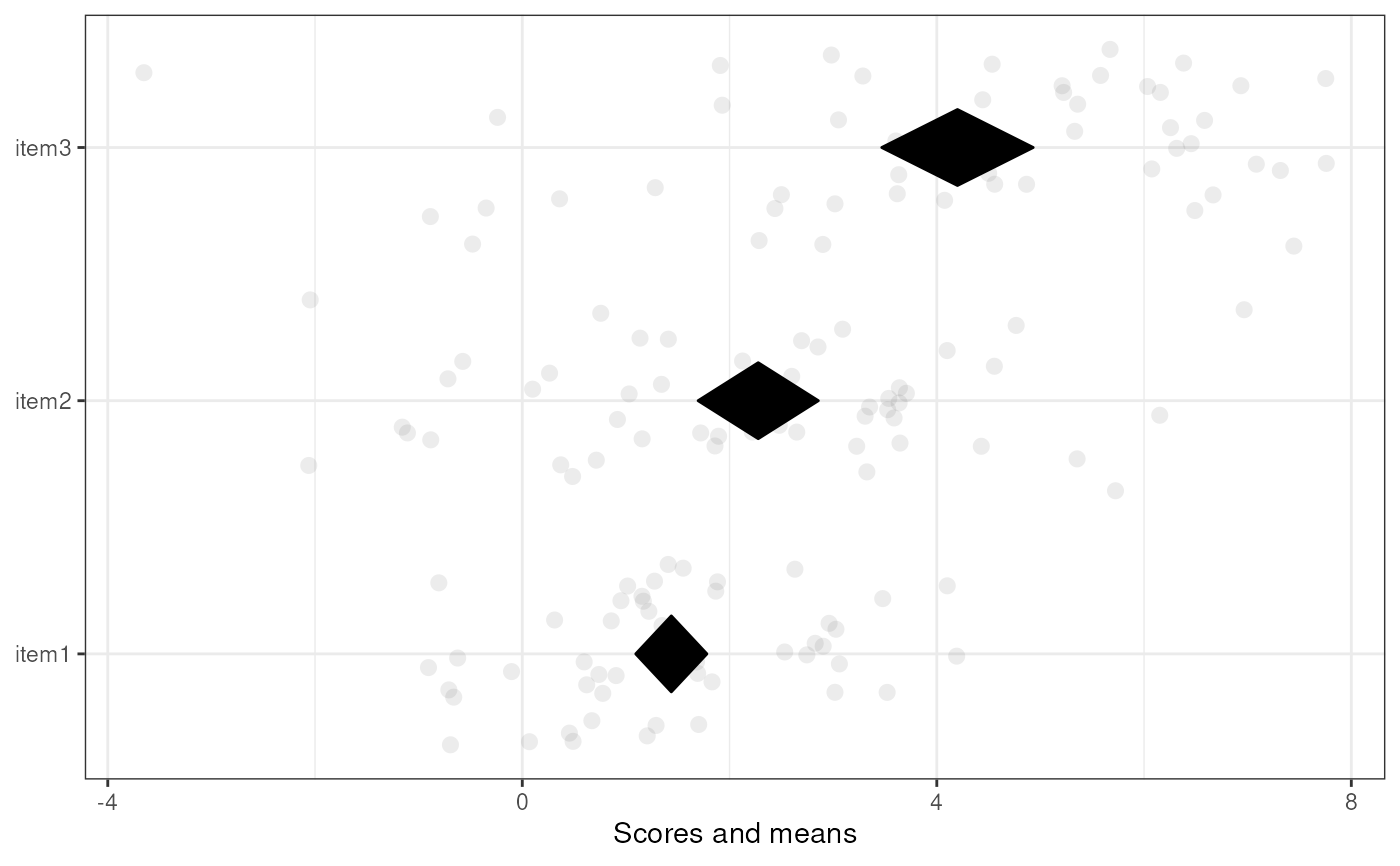
### A simple diamond plot
meansDiamondPlot(tmpDf);
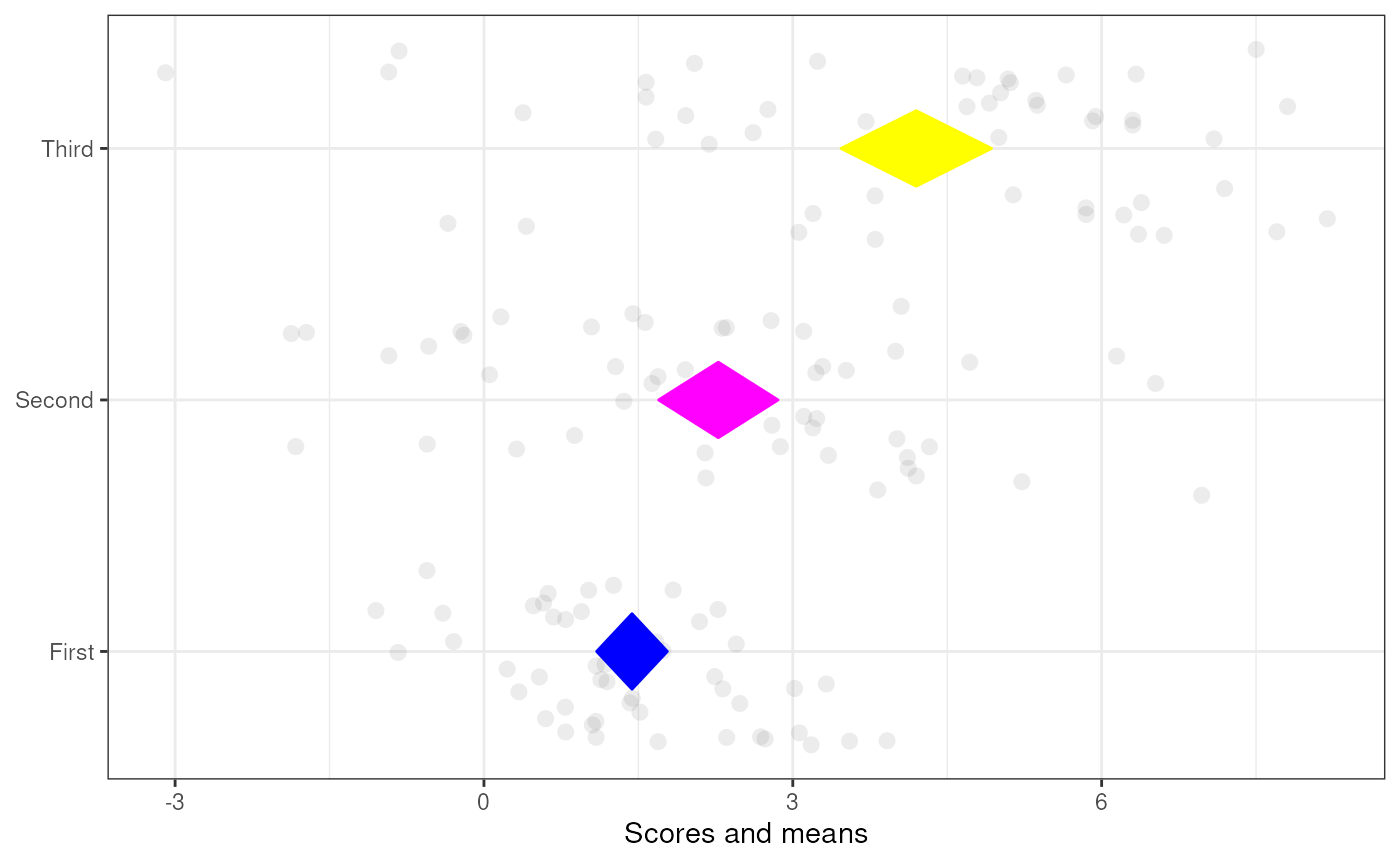
 ### A diamond plot with manually
### specified labels and colors
meansDiamondPlot(tmpDf,
labels=c('First',
'Second',
'Third'),
diamondColors=c('blue', 'magenta', 'yellow'));
### A diamond plot with manually
### specified labels and colors
meansDiamondPlot(tmpDf,
labels=c('First',
'Second',
'Third'),
diamondColors=c('blue', 'magenta', 'yellow'));
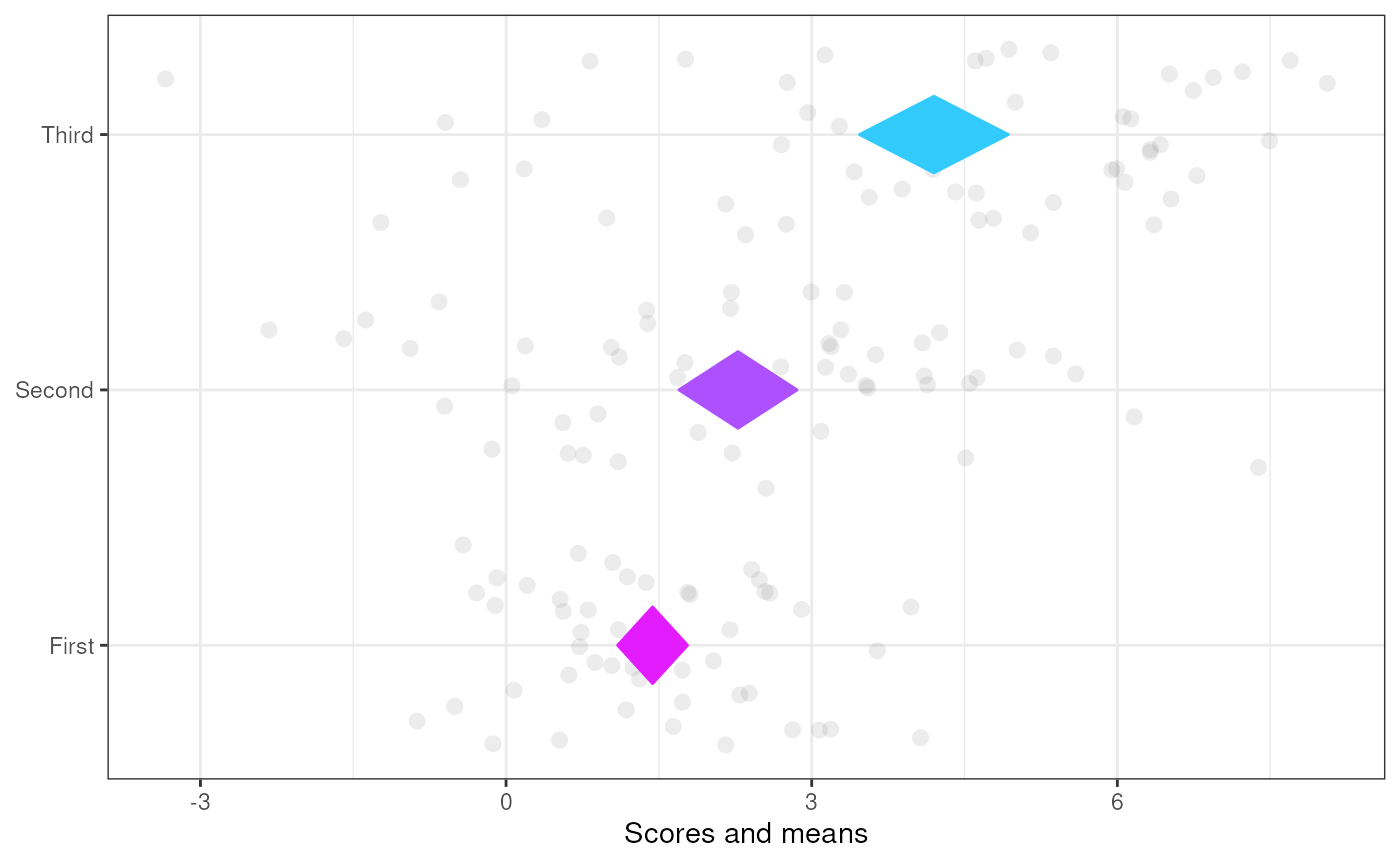
 ### Using a gradient for the colors
meansDiamondPlot(tmpDf,
labels=c('First',
'Second',
'Third'),
generateColors = c("magenta", "cyan"),
fullColorRange = c(1,5));
### Using a gradient for the colors
meansDiamondPlot(tmpDf,
labels=c('First',
'Second',
'Third'),
generateColors = c("magenta", "cyan"),
fullColorRange = c(1,5));